New drugs approved in the United States, by designations
What you should know about this indicator
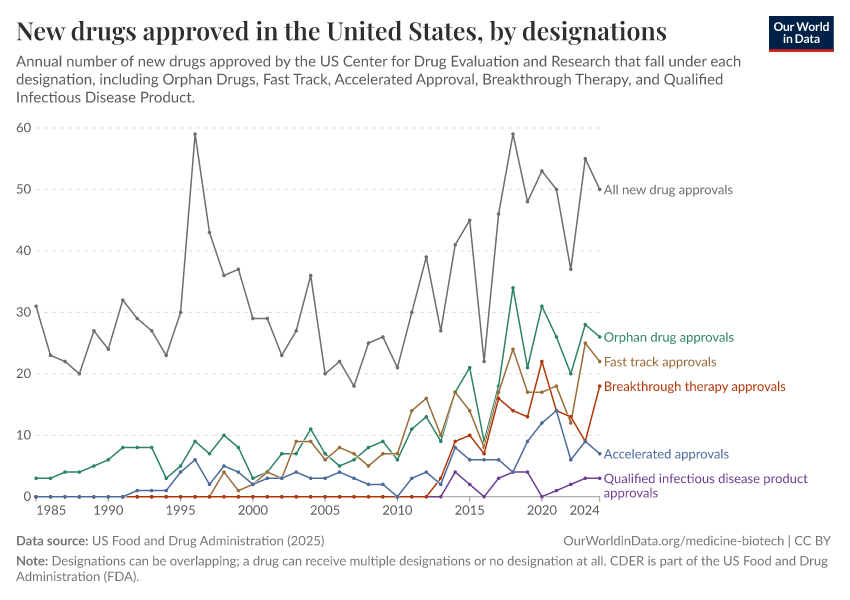
- This data only includes drugs approved by the Center for Drug Evaluation and Research (CDER). It does not include any biologics approved by the Center for Biologics Evaluation and Research (CBER) such as vaccines, allergenic products, or blood and blood components.
- Orphan drugs are drugs that treat rare diseases or conditions, affecting fewer than 200,000 people in the US. As developing these drugs would not be profitable for drug companies on their own, the Orphan Drug Act provides incentives such as tax credits and market exclusivity for these drugs.
- Fast track is a process created by the FDA to help drugs for serious conditions reach patients more quickly when there is an unmet medical need. It allows drug developers to have more frequent meetings with the FDA and to submit parts of their application as they’re ready, instead of waiting until everything is complete.
- Accelerated Approval is an FDA pathway that allows certain drugs for serious conditions to be approved based on earlier evidence — such as lab results or imaging — instead of waiting for direct proof of clinical benefit.
- Breakthrough Therapy is a designation the FDA can give drugs when early clinical evidence suggests that a new treatment may offer substantial improvement over existing options for a serious illness.
- A Qualified Infectious Disease Product is a drug intended to treat serious or life-threatening bacterial or fungal infections. The FDA grants this status under a law called the GAIN Act to encourage new antibiotics and antifungals.
- Designations can be overlapping; a drug can receive multiple designations or no designation at all.
- The data does not include generics or reformulations of existing drugs.
More Data on Medicine & Biotechnology
Sources and processing
This data is based on the following sources
How we process data at Our World in Data
All data and visualizations on Our World in Data rely on data sourced from one or several original data providers. Preparing this original data involves several processing steps. Depending on the data, this can include standardizing country names and world region definitions, converting units, calculating derived indicators such as per capita measures, as well as adding or adapting metadata such as the name or the description given to an indicator.
At the link below you can find a detailed description of the structure of our data pipeline, including links to all the code used to prepare data across Our World in Data.
Reuse this work
Citations
How to cite this page
To cite this page overall, including any descriptions, FAQs or explanations of the data authored by Our World in Data, please use the following citation:
“Data Page: New drugs approved in the United States, by designations”, part of the following publication: Tuna Acisu, Saloni Dattani, Fiona Spooner, Veronika Samborska, Hannah Ritchie, and Max Roser (2025) - “Medicine and Biotechnology”. Data adapted from US Food and Drug Administration. Retrieved from https://data-childlabor-ilounicef.owid.pages.dev:8789/20260318-114526/grapher/new-drugs-approved-in-the-united-states-by-designations.html [online resource] (archived on March 18, 2026).How to cite this data
In-line citationIf you have limited space (e.g. in data visualizations), you can use this abbreviated in-line citation:
US Food and Drug Administration (2025) – with major processing by Our World in DataFull citation
US Food and Drug Administration (2025) – with major processing by Our World in Data. “New drugs approved in the United States, by designations” [dataset]. US Food and Drug Administration, “CDER New Molecular Entity (NME) Drug and New Biologic Approvals” [original data]. Retrieved April 14, 2026 from https://data-childlabor-ilounicef.owid.pages.dev:8789/20260318-114526/grapher/new-drugs-approved-in-the-united-states-by-designations.html (archived on March 18, 2026).Download
Quick download
Download the data shown in this chart as a ZIP file containing a CSV file, metadata in JSON format, and a README. The CSV file can be opened in Excel, Google Sheets, and other data analysis tools.
Data API
Use these URLs to programmatically access this chart's data and configure your requests with the options below. Our documentation provides more information on how to use the API, and you can find a few code examples below.
Data URL (CSV format)
https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.csv?v=1&csvType=full&useColumnShortNames=falseMetadata URL (JSON format)
https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.metadata.json?v=1&csvType=full&useColumnShortNames=falseExcel / Google Sheets
=IMPORTDATA("https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.csv?v=1&csvType=full&useColumnShortNames=false")Python with Pandas
import pandas as pd
import requests
# Fetch the data.
df = pd.read_csv("https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.csv?v=1&csvType=full&useColumnShortNames=false", storage_options = {'User-Agent': 'Our World In Data data fetch/1.0'})
# Fetch the metadata
metadata = requests.get("https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.metadata.json?v=1&csvType=full&useColumnShortNames=false").json()R
library(jsonlite)
# Fetch the data
df <- read.csv("https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.csv?v=1&csvType=full&useColumnShortNames=false")
# Fetch the metadata
metadata <- fromJSON("https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.metadata.json?v=1&csvType=full&useColumnShortNames=false")Stata
import delimited "https://data-childlabor-ilounicef.owid.pages.dev/grapher/new-drugs-approved-in-the-united-states-by-designations.csv?v=1&csvType=full&useColumnShortNames=false", encoding("utf-8") clear